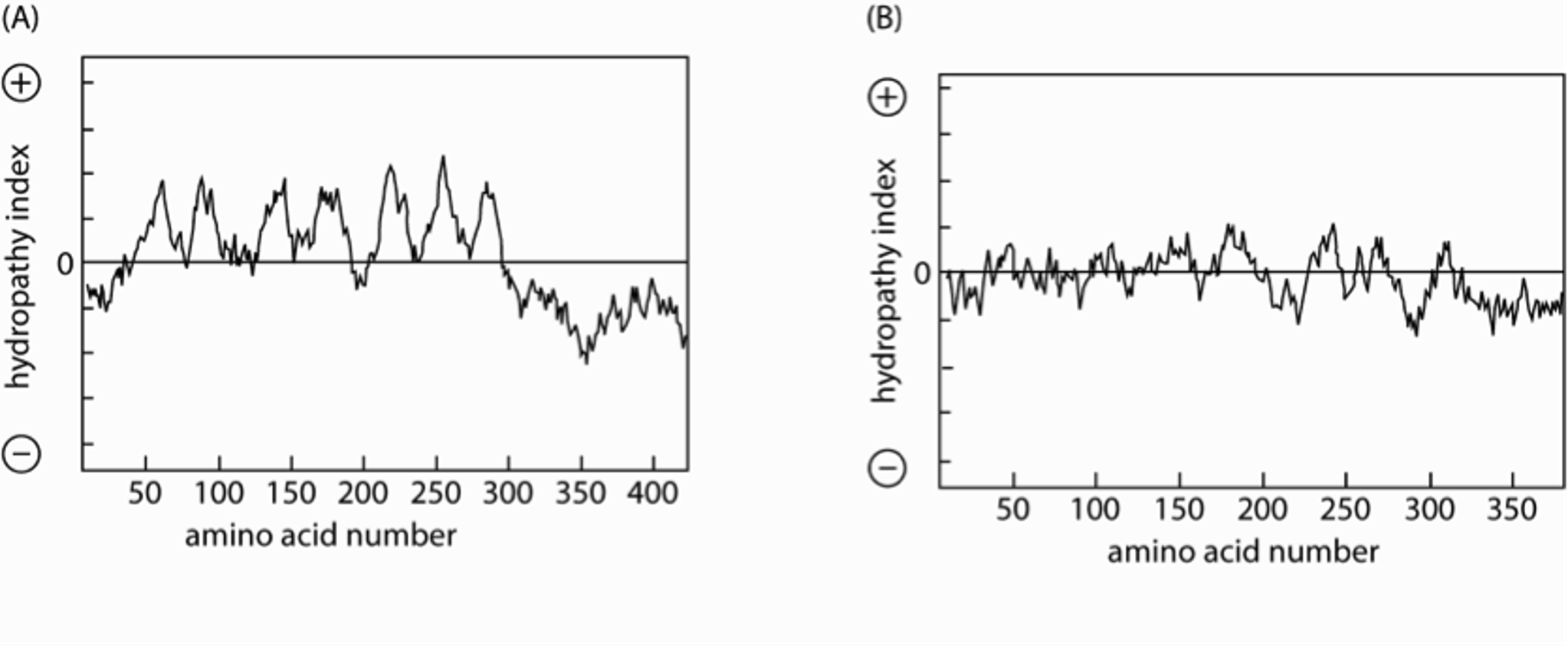
Analysis of signal sequence hydrophobicity. (A) Using the algorithm... | Download Scientific Diagram
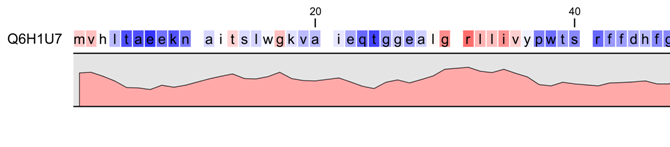
RNase k-02 hydrophobicity plot and Western blot analysis of HEK-293... | Download Scientific Diagram
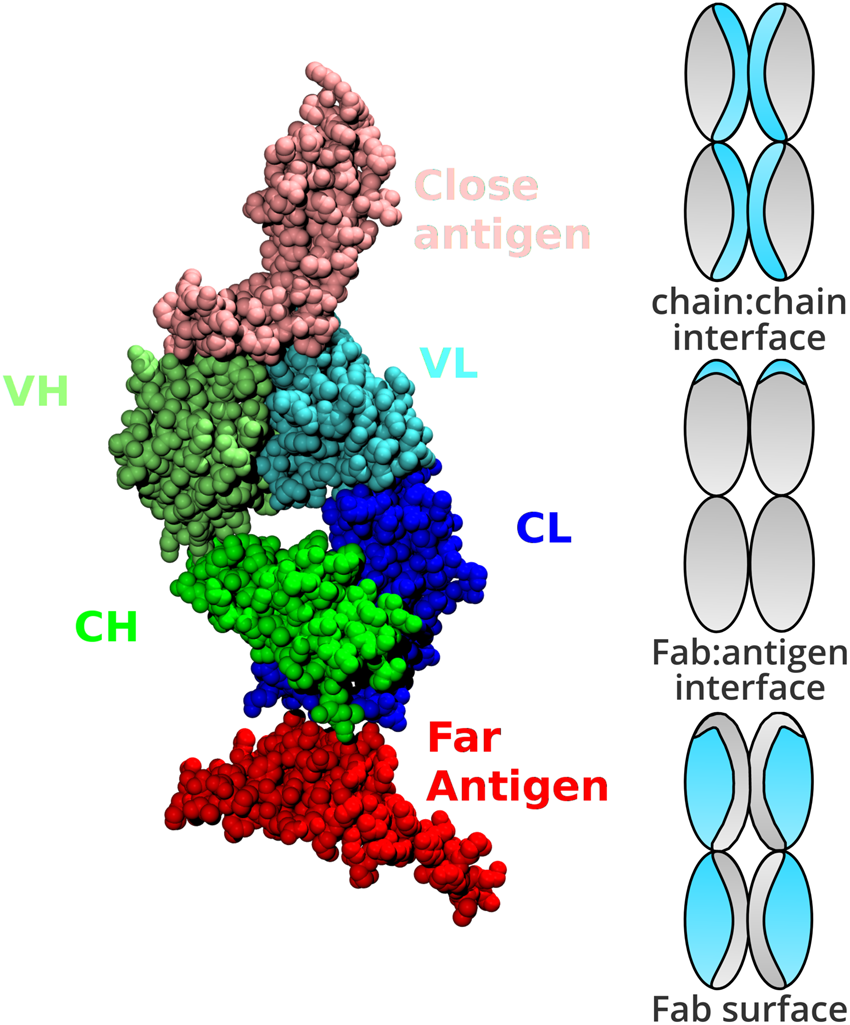
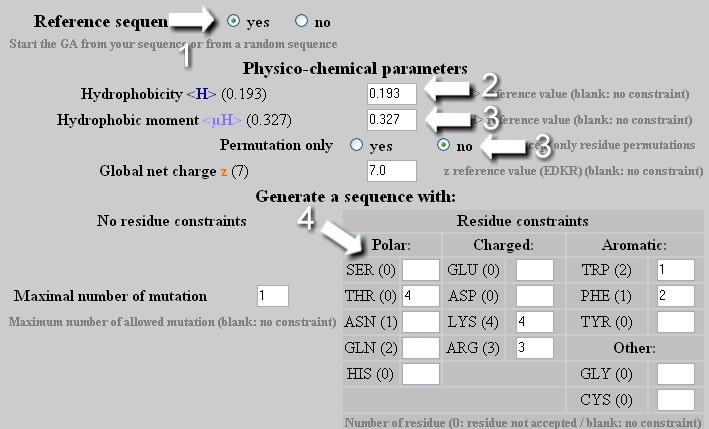
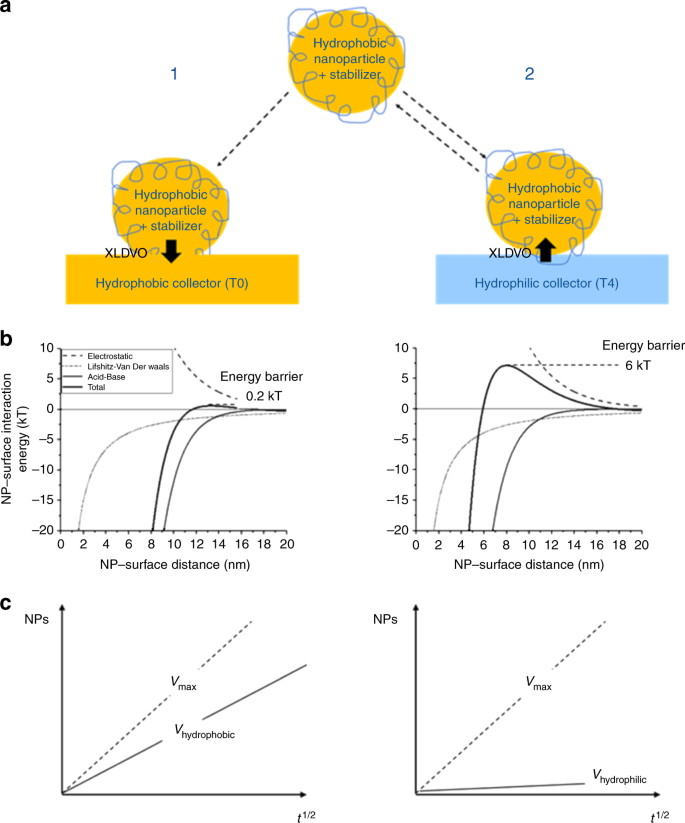
Web-based display of protein surface and pH-dependent properties for assessing the developability of biotherapeutics | Scientific Reports
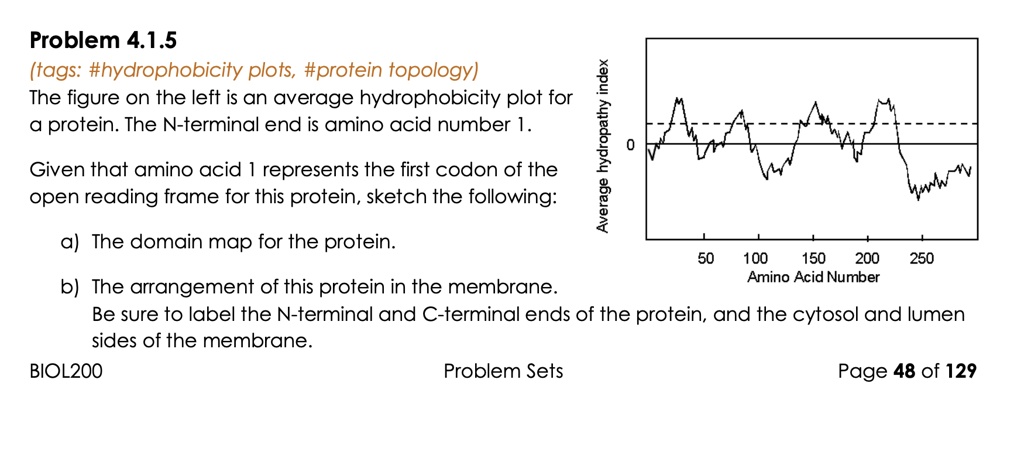
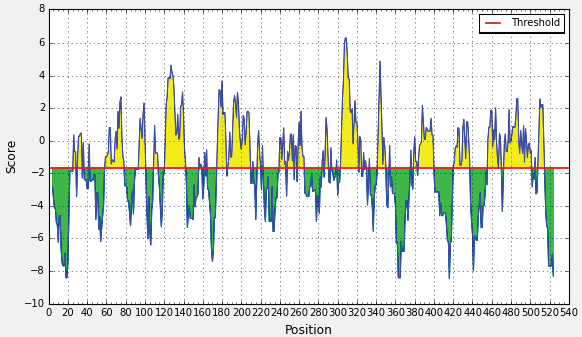
SOLVED: Problem 4.1.5 (tags: #hydrophobicity plots, #protein topology) 1 The figure on the left is an average hydrophobicity plot for protein. The N-terminal end is amino acid number 1 0 Given that

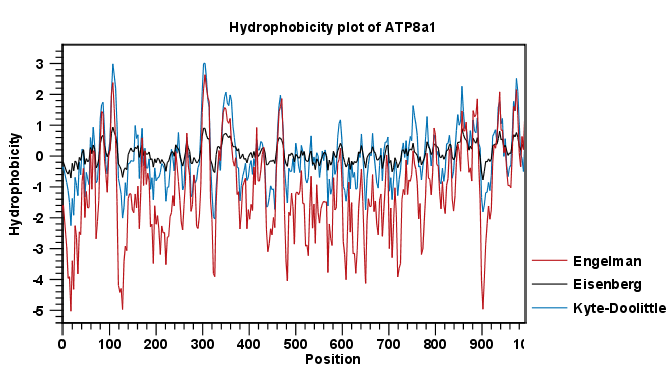
Methods of calculating protein hydrophobicity and their application in developing correlations to predict hydrophobic interaction chromatography retention. | Semantic Scholar
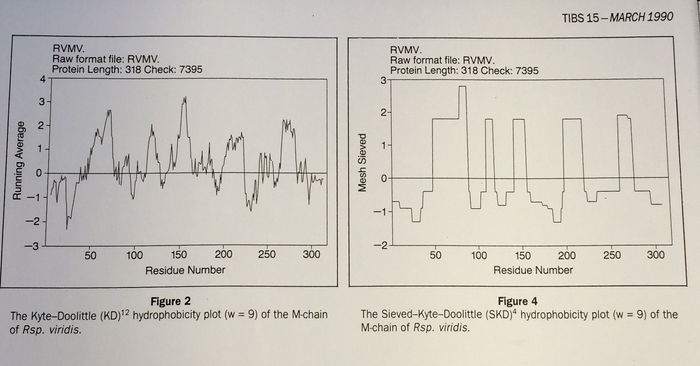
![PDF] Membrane protein structure prediction: Hydrophobicity analysis and the positive-inside rule | Semantic Scholar PDF] Membrane protein structure prediction: Hydrophobicity analysis and the positive-inside rule | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/0119a91510ebad911398166476f781fcecd73a6e/6-Figure4-1.png)
PDF] Membrane protein structure prediction: Hydrophobicity analysis and the positive-inside rule | Semantic Scholar

Methods of calculating protein hydrophobicity and their application in developing correlations to predict hydrophobic interaction chromatography retention - ScienceDirect